|
Perform scRNA data processing, cell type identification and differential expression analysis of cell clusters.Perform reference guided and de novo assembly of transcript sequences.Perform differential expression analysis.Estimate known gene and transcript expression.Align RNA-seq data to a reference genome. 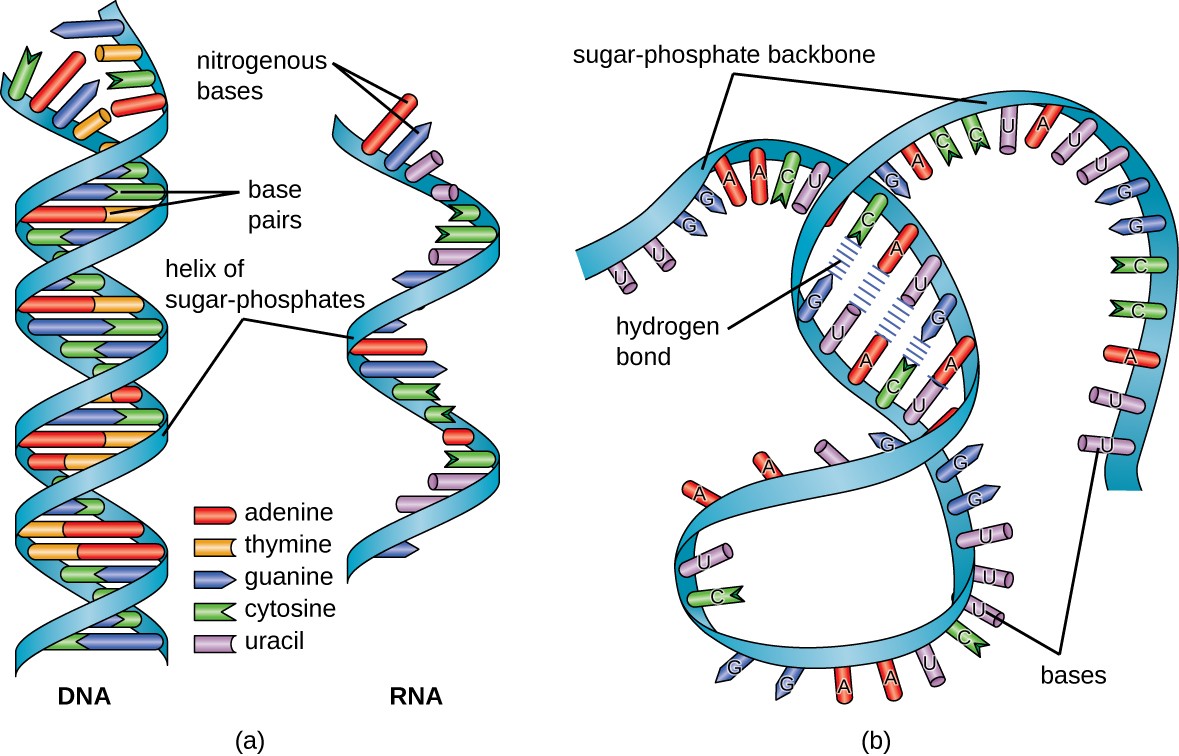
Perform command-line Linux based analysis on the cloud.Participants will gain practical experience and skills to be able to: The tutorials are designed as self-contained units that include example data (Illumina paired-end RNA-seq data) and detailed instructions for installation of all required bioinformatics tools (HISAT, StringTie, Kallisto, etc.). We have therefore developed this course to provide an introduction to RNA-seq and scRNA-seq data analysis concepts followed by integrated tutorials demonstrating the use of popular bioinformatics analysis packages. Similarly single cell RNA sequencing (scRNA-seq) has been widely adopted for experimental systems where it is crucial to understand a complex mixture of cell types. RNA sequencing (RNA-seq) has rapidly become the assay of choice for robustly interrogating RNA transcript abundance and diversity. 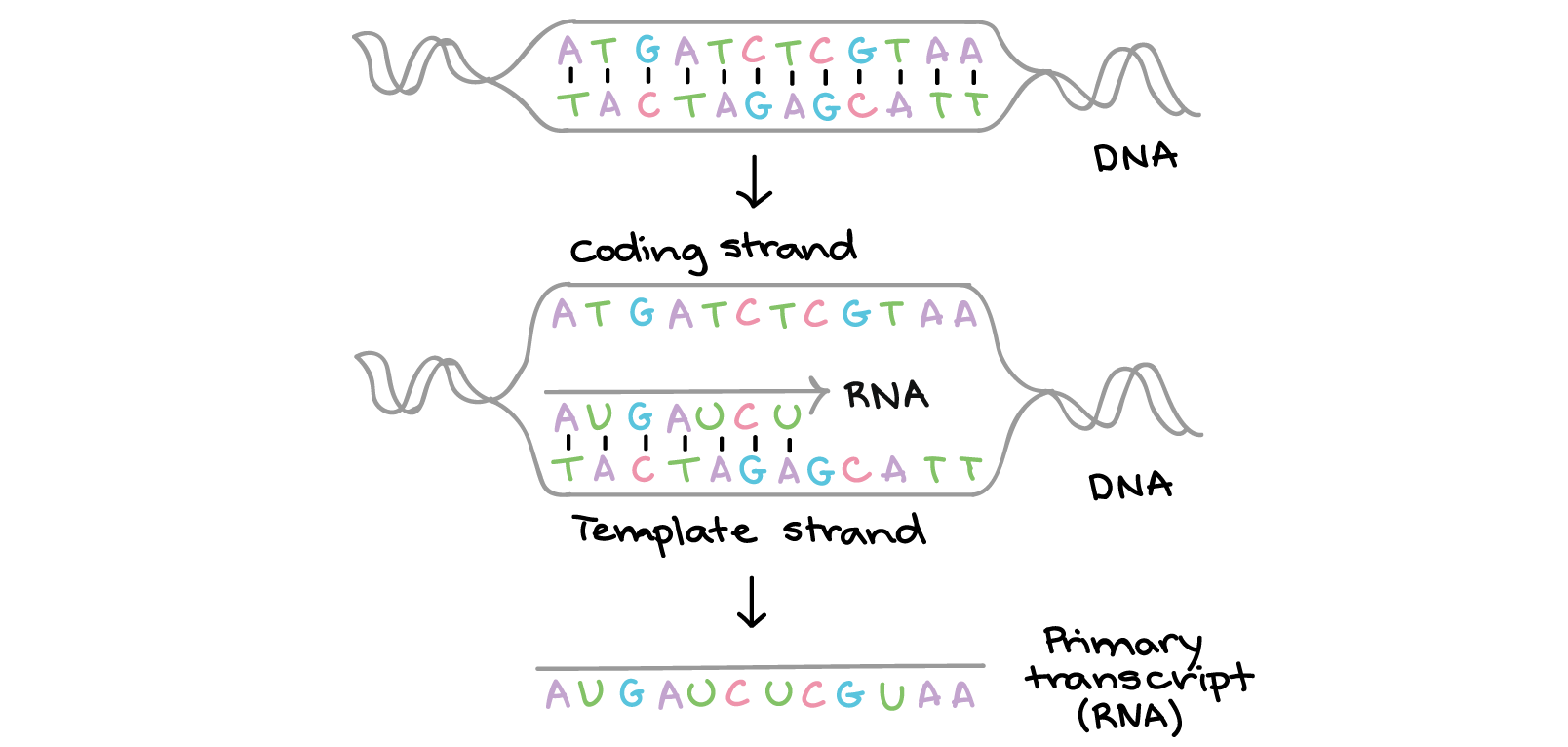
The age of affordable massively parallel sequencing has exponentially increased the availability of transcriptome profiling. Each page has a course link at the top to bring you back to the table of contents. 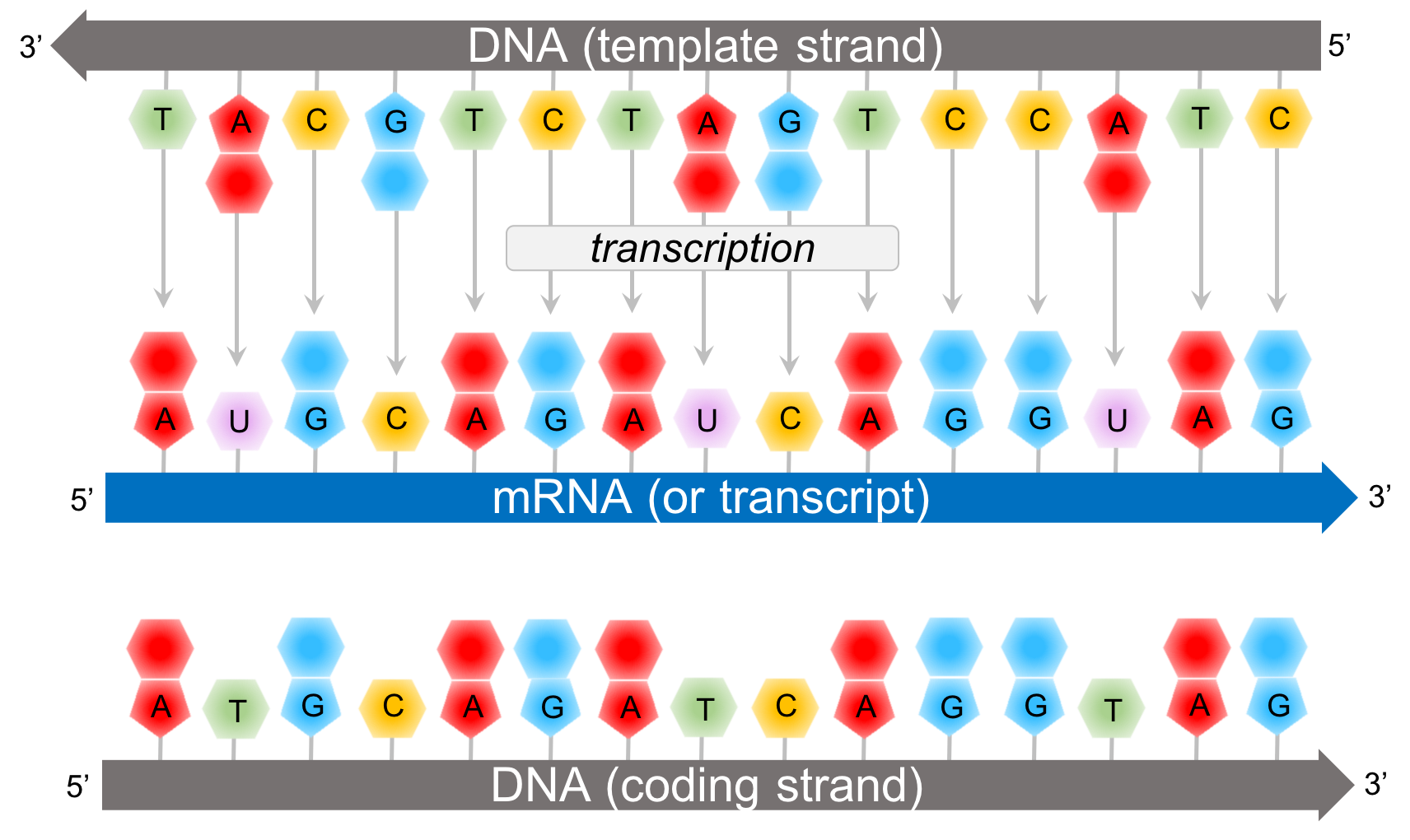
Use the course page to navigate your way through all exercises. Welcome to the RNA-seq Bioinformatics Course. Introduction to bioinformatics for RNA sequence analysis RNA-seq
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
